The Work
A private research institute in a laptop.
Three months. One M5 Max. Three engines.
Source: github.com/norayr-m
A private research institute in a laptop.
Three months. One M5 Max. Three engines.
Source: github.com/norayr-m
These six are what the work is. The next six pages introduce each pillar in one breath. Then we go in.
A ternary lattice running on Apple Silicon GPU, scheduling cells in seven phases via distance-2 chromatic separation on a hexagonal grid. The substrate that hits the central GCUPS number on a laptop.
at 1 billion hex cells · 522 ms/tick · M5 Max · 10-run published, 95% CI
Read more: Savanna Engine repo · Savanna white paper · 7-coloring explained — interactive
Every node holds a 64-bit truth table — six inputs, one output, the same primitive that lives at the heart of an FPGA. Nodes wired into a ranked acyclic graph; the engine evaluates the whole graph in parallel on the GPU each tick. Snapshots, write-ahead log, multi-version reads — durable like a real database.
The BACK_EDGE primitive lets state latch across tick boundaries — unlocks AC-3 constraint propagation, Hopfield-style recurrent networks, Boolean networks with feedback.
Read more: DagDB repo · DagDB white paper · Engine site · SQL architecture
Geometric reasoning over protein conformations. Substrate, not folding prediction. AlphaFold gives you the structure; we operate on it.
The hepatocyte 10⁷ Pass 2 schema lays in a spatial-adjacency layer with hepatic-zonation policy ranks (sixteen levels, periportal to pericentral). The graph encodes what is adjacent in space; the engine reasons over the topology.
Read more: Isomorphic Walk repo
Memory layout via Morton Z-curve — cache-friendly traversal that delivers a 2.11× speedup over row-major at one billion cells. Distance-2 chromatic separation on the hexagonal grid uses exactly seven colours (Molloy–Salavatipour 2005), with the engine using (col + row + 4·(col & 1)) mod 7 as the colour-class formula. The combination is what makes the substrate parallelizable on commodity GPU hardware.
Read more: Savanna Morton charts · 7-coloring explained
Apple unified memory used as zero-copy substrate — no CPU/GPU buffer boundary. Tile-streaming scaffold for 10¹¹ nodes on one M5 Max: page tiles to and from NVMe, two tiles resident at a time, cross-tile addressing via 24-bit-tile + 40-bit-local u64 global ID.
Three-environment separation — dev / test / prod under guarded paths. Triple-write to markdown, DagDB, and Postgres with markdown as the leader.
Read more: DagDB engine repo · DagDB white paper
The substrate the simulation engines run on assumes regularity properties of the underlying space. CVPDE-class results — Almgren-monotonicity-style — are foundation lemmas that have to hold for the substrate to behave the way the engineering claims.
Open directions: extension to non-uniform grids, generalization beyond hexagonal lattices, sharper bounds on the chromatic number under non-uniform-distance constraints.
Read more: DagDB engine repo
The 100-billion-cell existence proof — and where the M5 thermal ceiling actually lives.
Calibration: this slide reports a VERIFIED build. Savanna ran 100 billion cells over 9 hours on one M5 Max laptop, peaked 15.8 GCUPS, wrote 500 GB of state to disk. Same dispatch + Morton + 7-coloring machinery DagDB inherits — making this the existence proof for DagDB’s trillion-cell scale path. Honest framing guard: Savanna’s FLUID kernels (scent, density, predator-prey, peristaltic streaming) are intentionally NOT in DagDB. Conflating the two is overclaim risk. See
one_pager.md’s “DagDB inherits Savanna’s physics engine” overclaim callout for the precise lineage boundary.
A predator-prey-grass spatial-lattice simulator on Apple Metal GPU, running discrete biology on a 1024×1024 hex grid as a test workload for the underlying spatial-compute machinery. The biology is the load — the engine is what’s being measured.
Three engineering primitives load-bearing the throughput:
Same machinery DagDB inherits (HexGrid, Carlos Delta encoder, halo file format). Different per-cell state: Savanna stores fluid + scent + entity tables; DagDB stores Boolean LUT6 + ranked DAG metadata.
Existence proof for the trillion-cell scale path.
The verified 100 B / 9 h Savanna run on one M5 Max laptop is the highest
scale the substrate machinery has actually shipped. DagDB’s
tile-streaming spec for 10¹¹ nodes
(docs/tiled-streaming.md) targets the same hardware
envelope. Without Savanna’s existence proof, DagDB’s trillion-cell goal
is just a target. With it, the engineering question reduces to “can we
tile the LUT6 / BACK_EDGE state across NVMe at the same machinery’s
throughput envelope?” — a tractable engineering question rather tha
feasibility one.
| Grid | Throughput | Wall | State | Real-time? |
|---|---|---|---|---|
| 1 M cells | 1 634 tps | sub-second | 23 MB | ✓ |
| 16 M cells | 77 tps | ~13 s | 367 MB | ✓ |
| 64 M cells | 19 tps | ~52 s | 1.5 GB | ✓ |
| 1 B cells | 1.2 tps | ~14 min | 23 GB | ✓ |
| 100 B cells | ~3 tps* | 9 hours | 500 GB | (thermal-throttled) |
Peak: 27.1 GCUPS measured at 16 M cells, 10-run validated, < 1 % standard deviation. Sustained 100 B: 15.8 GCUPS — figure cited on the deck.
Saw the predator-prey simulation collapse repeatedly to extinction: lions kill all zebras, then starve. Classic thermodynamic suicide switch. Gemini Deep Think’s diagnosis: the per-tick lion energy update needed a satiation cap so well-fed lions stop hunting and become a “physical meat shield” — preventing the runaway overhunting that triggered the death spiral.
Implementation: one inequality in the lion tick_phase
kernel (if my_energy >= SATIATION_THRESH: don't_hunt).
Predator-prey populations stabilize at oscillating limits. Same fix
Holling 1959 prescribes for ODE-level Lotka-Volterra; we just learned
why it’s load-bearing on a discrete lattice the hard way.
The white paper’s eight-overcome teardown lists the full set — including the LCG-parity “ghost wind” hash bug, the chromatic- advection dispatch-order drift, the Atto-Fox sub-individual extinction trap, and the Voronoi-formation “lion haboob” that emerged unbidden from the corrected dynamics.
dagdb/Sources/DagDB/HexGrid.swift (lifted verbatim),
Sources/DagDB/CarlosDelta.swift (encoder preserved).
Per-cell state and biology kernels NOT inherited.| Claim | Label |
|---|---|
| 100B-cell run, 9 hours, 15.8 GCUPS | VERIFIED |
| 7-coloring proves topological cleanness (no Euler scars) | VERIFIED |
| Morton Z-curve 2.11× at 1 B cells | VERIFIED |
| Type II satiation prescribed by Deep Think; broke the death spiral | VERIFIED (caught + fixed; whitepaper §4) |
| Same dispatch + Morton + 7-coloring machinery as DagDB | VERIFIED (code-level lineage) |
| Savanna’s fluid kernels run on DagDB | OVERCLAIM — DO NOT SAY |
| 100 B cell + DagDB LUT6 = trillion-cell DagDB | SPEC (engineering target; tile-streaming spec’d not built) |
What’s not claimed. Savanna is a fluid-dynamics-capable spatial lattice engine — scent diffusion, peristaltic streaming, predator- prey kinetics. DagDB is not. The lineage between them is dispatch patterns + halo file format + Morton + 7-coloring. The fluid / scent / density kernels were intentionally dropped when DagDB became Boolean-LUT-only. Conflating the two = the precise overclaim risk flagged in
one_pager.md. The existence proof Savanna provides for the scale envelope is real; the engineering question for DagDB at 10¹² is “can we tile LUT6 + BACK_EDGE state across NVMe at this throughput envelope” — which is a tractable port, not a feasibility claim.
Filled by slvr 2026-05-01. Pub-screen gate before commit to assets.
A graph where every node carries its own program — and the whole graph runs in parallel each GPU tick.
Owner: dag (engine internals); pub (editorial pass)
Status: DRAFT 1 (2026-05-02) Tier: 2
(medium level / main directions) Calibration anchor:
../one_pager.md
One-slide condensation of DagDB’s engine story, sized for the architect-grade audience to grasp the engine class and what makes it different.
Required content: - Headline: graph database where
every node is a 64-bit truth table (LUT6); ranked DAG, evaluated on the
GPU each tick - BACK_EDGE primitive (synchronous-circuit register
pattern) verified via AC-3 round-trip — VERIFIED, 120+
Swift tests green - Snapshot v4 + WAL + MVCC — durable like a real
database - Bitwise LUT composition (COMPOSE AND/OR/XOR/NOT)
for runtime graph simplification - ~2 ms per tick at 1 M nodes on M5 Max
(engine throughput at the demo scale) - Pointer to the engine repo + the
wiki tab at github.com/norayr-m/dagdb-engine/wiki
What this slide does NOT do: - Claim “thinking
engine” — explicitly off per one_pager.md. Speedboat for
one narrow class of structured-dependency reasoning. - Mix
Savanna-physics framing in (those are different engines)
Headline. A graph database where every node holds a 64-bit truth table — i.e., a programmable Boolean function with up to six inputs. Same primitive that lives at the heart of an FPGA. Nodes wire into a ranked acyclic graph; the engine evaluates the whole graph in parallel on the GPU each tick. Persistent like a real database — snapshots, write-ahead log, multi-version reads. One machine.
The four verified pieces.
rank(src) > rank(dst)) makes topological order
well-defined; LUT6 means each gate is one indirect read of a 64-bit
integer.reg / non-blocking assignments. Verified end-to-end by AC-3
Australia 3-coloring round-trip against an independent Python reference:
per-tick equality across 21 register nodes, converges in 2 synchronous
ticks. 120+ Swift tests green. Without BACK_EDGE the
substrate hosts only feed-forward Boolean DAGs; with it, AC-style
constraint propagation, Hopfield-shape recurrent networks, and any
register-clocked dynamical system become natively expressible.F_FULLFSYNC + replaceItemAt + dir fsync
(Apple-SSD discipline). WAL replay rolls forward through a crash. Reader
sessions take snapshots without blocking writers — concurrent queries
see a consistent view of the graph.COMPOSE AND src1 src2 INTO dst, plus
OR/XOR/NOT. Collapses subgraphs into single LUTs without
re-evaluating the original tree at runtime. Useful for graph
simplification passes and for synthesizing complex gates from
primitives.Throughput. ~2 ms per tick at one million nodes on M5 Max.
Lifted verbatim from
dagdb-engine/site/sql-architecture.html (slide 15).
Surface. Engine repo
github.com/norayr-m/dagdb-engine, wiki tab on the same
repo, browser-runnable demos at
norayr-m.github.io/dagdb-engine/site/. The BACK_EDGE wiki
page is the deepest written reference for the primitive.
https://github.com/norayr-m/dagdb-engine/wiki/Back-edgeshttps://github.com/norayr-m/dagdb-enginehttps://norayr-m.github.io/dagdb-engine/site/index.html../one_pager.md: BACK_EDGE/AC-3,
Snapshot v4, MVCC, COMPOSE LUT — all VERIFIED| Claim | Label |
|---|---|
| 6-bounded ranked DAG with LUT6 gates | VERIFIED |
| BACK_EDGE primitive, AC-3 verified | VERIFIED — 120+ tests, per-tick equality with Python reference |
| Snapshot v4 + WAL + MVCC | VERIFIED |
| Bitwise LUT composition (COMPOSE) | VERIFIED |
| ~2 ms per tick at 1 M nodes on M5 Max | VERIFIED — measured |
| Substrate hosts AC-style iteration to fixed point | VERIFIED via the AC-3 round-trip; substrate-class generalisation NOT claimed |
| 10¹¹ tile-streaming on one M5 | SPEC — Step 4 scaffold landed, Steps 5–7 pending |
What’s not claimed. DagDB is not a general thinking engine. Missing: variables, quantifiers, first-order logic, probabilistic truth, search/planning, recursive within-tick evaluation. It is a fast specialised substrate for one class of structured-dependency reasoning.
Stubbed by Pub on 2026-05-01. Drafted by dag on 2026-05-02. Pub-screen pass before any external recording or distribution.
Source: github.com/norayr-m/dagdb-engine · wiki · demos
Asking the substrate where, inside a protein, distant binding sites can talk to each other.
Owner: fold (biology/math content); pub (editorial
pass) Status: DRAFT 1 Tier: 2 (medium
level / main directions) Calibration anchor:
../one_pager.md
A method for predicting allosteric pathways in proteins from a single static structure (the kind crystallographers deposit in the PDB). The method treats residues as nodes in a graph and 3D contacts as edges, then runs fast graph algorithms — breadth-first search and sparse spectral methods — to identify the corridor of residues that mediates signal between two designated functional sites.
Implemented as a Swift CSR breadth-first-search primitive on Apple
Silicon, exposed to Python via ctypes for analysis
pipelines. Two findings cleared pre-registered acceptance gates with
Gemini 2.5 DeepThink as neutral arbiter; one
algorithmic variant was rejected under the same protocol; two follow-up
variants ran as pilots that closed without warranting formal
pre-registration. All numbers below were locked before data
collection.
Allostery is one of the central mechanisms of biological regulation — GPCRs that account for roughly a third of clinically relevant drug targets, ATP-driven motors, kinase signaling cascades, hemoglobin’s cooperative binding. Identifying which residues actually carry the signal between two functional sites is hard because the residues that physically lie between them are usually a much larger set than the residues that biologically matter.
Standard computational tools for this question fall into two categories. Molecular dynamics simulates atom-by-atom under realistic physics — gold standard, but a microsecond on a million- atom system takes days on a GPU cluster. Elastic Network Models (GNM, ANM) treat residue contacts as springs and solve a matrix eigenvalue problem — cheap but O(V2) memory and O(V3) time; the dense Kirchhoff matrix doesn’t fit at viral-capsid scale. This work explores a third path: discrete graph algorithms, O(V + E) sparse, that run on a laptop.
Asymptotic scaling. On the 313,236-atom HIV-1
capsid (PDB 3J3Q), the Swift BFS finishes a full traversal in 6
milliseconds on M5 Max, sweeping 4.27 M contact edges.
Sustained throughput ~250 million traversed edges per second, 4 ns per
edge. ProDy GNM cannot run on this system: dense Kirchhoff requires
313, 2362 × 8 B ≈ 784 GB of
memory. This is a memory-exclusion claim, not a wall-clock
comparison — the two tools answer different questions; the
comparison only tells you which question is feasible at a given scale.
Reproducible at
experiments/2026-04-20_scaling_vs_prody/scaling_v2.py.
Allosteric pathway recovery — ESTABLISHED under pre-registered protocol. On a cohort of 10 well-studied allosteric systems (IGPS, PTP1B, β₂-AR, Hsp70, PKA, GlmS, ATCase, tryptophan synthase, PFK, Abl), a BFS suboptimal-tube on an 8 Å Cα contact graph identifies published allosteric-pathway residues better than a Euclidean-cylinder baseline in 6 of 7 evaluated systems. 3 systems were pipeline failures (seed-residue parsing) and were dropped without substitution per the locked protocol — no retroactive cohort manipulation. Median ΔF1 = +0.0707. Pre-registered acceptance gate (locked with the arbiter before any data were collected): ΔF1 ≥ +0.05 AND ≥ 70% wins. Both cleared. Ground-truth residue lists from primary literature (Rivalta 2012 / Wiesmann 2004 / Venkatakrishnan 2013 / Zhuravleva 2012 / Taylor 2012 / Lipscomb 2008 / Schirmer & Evans 1990 / Azam 2008).
Fiedler zero-crossing. Standalone variant: 3 of 9 evaluated wins versus BFS-tube, median ΔF1 = −0.020. Intersection variant (BFS-tube ∩ Fiedler-ZC): 4 of 9 wins, median ΔF1 = −0.012, with two large per-system wins (PTP1B +0.165, PFK +0.101) offset by one structurally-meaningful catastrophic loss (Abl, where the bend axis and the BFS path between A-loop and αC-helix are graph- disjoint, producing zero intersection). Conclusion: standalone ZC not worth formal pre-registration; intersection variant is precision-rich but recall-poor, parked pending a precision-at- fixed-recall evaluation framework.
λ₃ multi-mode spectral subspace. The arbiter named “λ₂ through λ₅ subspace” as a possible recovery direction. Pilot tested the smallest extension toward that idea: λ₃ alongside λ₂ across three formulations (standalone, union, 2D subspace axis). All three produced median ΔF1 ≈ −0.17, 2 of 9 wins each. Adding v₃ to the borderline intersection variant dilutes it (median ΔF1 dropped from −0.012 to −0.031). Multi-mode subspace direction closed without consuming a formal pre-registration round. Walking out to λ₄–λ₅ is unwarranted: pattern is “v₃ already adds noise faster than signal.”
Swift package: BFSLib (C-callable dylib) +
KowalskiCrush (CLI) + KowalskiCrushGPU (Metal
compute, with measured-13×-slower-than-CPU caveat documented in source
for single-source BFS at biomolecule scale; would win for multi-source
dispatch or graphs ≥ 10⁶ nodes). Python ctypes wrapper, zero-copy via
Apple Silicon unified memory.
| Claim | Label |
|---|---|
| HIV-1 capsid 313,236 atoms, 6 ms BFS sweep | VERIFIED |
| 784 GB ProDy GNM memory exclusion | VERIFIED |
| Allosteric cohort 6 of 7 wins, median ΔF1 = +0.0707 | ESTABLISHED (pre-registered) |
| Spectral snap walk REJECTED | RECORDED (locked criteria) |
| Fiedler zero-crossing standalone pilot | PILOT (not pre-registered) |
| Fiedler zero-crossing intersection variant | PILOT (borderline, parked) |
| λ₃ multi-mode subspace pilot | PILOT (rejected the direction) |
../one_pager.md: HIV-1 capsid
313,236 atoms / 6 ms BFS sweep — VERIFIED; allosteric cohort 6 of 7
wins, median ΔF1 = +0.0707 — ESTABLISHED (pre-registered); spectral snap
walk REJECT — RECORDED (locked criteria)slides/hepatocyte_pass_2_spec.mdexperiments/2026-04-20_scaling_vs_prody/scaling_v2.py
(asymptotic scaling),
experiments/2026-04-20_allosteric_cohort_n10/cohort.py (the
cohort-establishing run),
experiments/2026-04-21_isowalk_spectral_snap/snap_walk.py
(the recorded REJECT)What’s not claimed. No validated bio digital twin. The IsoWalk substrate is verified at the engine level (BFS / spectral methods on biomolecular contact graphs); specific tissue-twin validation against published biology data is future work, not present work. The norm-growth inequality from earlier prose (Matevosyan + Petrosyan, in preparation) has been retracted in v0.1; references to it are part of the honesty story, not operative claims.
Drafted 2026-05-01 by fold per kickoff drop. Fill matches Dag’s one_pager.md three-column discipline.
The lineage itemized — the prior work the substrate leans on, and from whom.
Owner: ref (citations + lineage); pub (editorial
pass) Status: DRAFT 1 — populated 2026-05-01
Tier: 2 (medium level / main directions)
Calibration anchor: ../one_pager.md
Novel combination, prior-art ingredients. The applied-math machinery powering Savanna + DagDB + the trio is, at the level of individual ingredients, all classical or near-classical. The contribution is the assembly — a single-laptop spatial lattice engine with topological-cleanness, cache-line locality, ecologically-stable Lotka–Volterra, lossless compression, and CFL-safe temporal compression all stacked on one Apple Silicon GPU. The combination matters.
Where it appears. The Savanna and DagDB engines partition the hexagonal cell lattice into 7 color classes such that no two cells in the same class are within radius-1 (Moore-equivalent on hex). Same-color cells can be updated in parallel without read-write hazard — this is what makes lock-free chromatic dispatch possible on the GPU.
Why 7. On the hex lattice, the squared graph G2 has chromatic number exactly 7 — equivalently, every vertex has 6 distance-1 neighbours, and 7 colors saturate the bound by Brooks-style argument refined for planar bounded-degree graphs.
Citations. - Molloy & Reed (2002). Graph Colouring and the Probabilistic Method. Springer, ISBN 978-3-540-42139-4. The local-bound machinery for χ(G2) on bounded-degree planar graphs. - Salavatipour, M.R. (2005). The complexity of L(p, q)-labeling of planar graphs. Discrete Math. 285:227–240. Refines the planar bounded-degree case relevant to hex squared graph.
Label. VERIFIED prior art — engineering inherits a textbook bound.
Where it appears. Cell-state buffers in DagDB and Savanna are stored in Morton order (bit-interleaved (x, y) → linear index). At dispatch time, the 7 same-color cells in any tile are guaranteed to share L2 cache lines on M5 because Morton-adjacent cells map to Morton-adjacent linear indices.
Verified speedup. 2.11× at 109 cells over row-major layout — measured on M5 Max, reproducible. Number lives in the one-pager.
Citations. - Morton, G.M. (1966). A computer oriented geodetic data base and a new technique in file sequencing. IBM Tech. Report. Original Z-order curve. - Bader (2013). Space-filling curves: an introduction with applications in scientific computing. Springer. Modern reference; L2-cache-line alignment property at 2D 8-neighbour Hamming-ball is Theorem 4.x there.
Label. VERIFIED — measured on the substrate. Algorithmic locality is prior art; the speedup number is ours.
Where it appears. Savanna’s Lotka–Volterra-class predator–prey dynamics. The classical Lotka–Volterra (linear functional response) produces an unphysical predator death spiral at scale — predators eat faster than prey reproduce, populations crash to zero. Holling Type II’s saturating-rate kernel f(N) = aN/(1 + ahN) is the fix.
Citation. - Holling, C.S. (1959). Some characteristics of simple types of predation and parasitism. Canadian Entomologist 91(7):385–398. Original Type I / II / III taxonomy. Type II is the saturating one.
Label. VERIFIED prior art — engineering inherits the textbook correction.
Where it appears. Savanna 100B-cell state file at 9 hours hits the M5 thermal ceiling but produces a 500 GB raw state. Carlos Delta brings that to ~10 GB lossless via XOR + Zstandard.
Citation. - Mateo, C. Carlos Delta. MIT-licensed implementation of XOR-then- Zstandard on dense numeric arrays. Independently developed by Carlos Mateo (external collaborator). 50× lossless ratio measured on Savanna state files.
Label. VERIFIED external. Credit is Carlos Mateo’s; the 50× number is measured on our state files.
Where it appears. Savanna’s dt-compression schedule (skipping ticks when the simulation is in a slow regime) needs a safety clamp so signal information cannot propagate faster than the discrete grid resolves. CFL gives the clamp.
Citation. - Courant, R., Friedrichs, K., Lewy, H. (1928). Über die partiellen Differenzengleichungen der mathematischen Physik. Math. Ann. 100:32–74. The original CFL condition for hyperbolic PDEs; carries over cleanly to discrete CA dynamics with a maximum signal speed.
Label. VERIFIED prior art — engineering inherits the textbook condition.
DagDB is a novel combination, but each ingredient sits on visible prior art. Naming the lineage is honest framing:
| DagDB ingredient | Prior art | What’s the same | What’s different |
|---|---|---|---|
| LUT6-as-data | FPGAs (Xilinx, Altera) | 6-input lookup table as the primitive computational unit | DagDB stores LUTs as graph nodes and evaluates them on a CPU/GPU via Metal; FPGAs burn them into silicon |
| Ranked DAG with parallel evaluation | Pregel (Malewicz 2010), GraphLab (Low 2010, 2012) | Vertex-program graph evaluation under a synchronisation barrier | DagDB’s rank invariant (rank(src) > rank(dst)) gives a static evaluation order; Pregel/GraphLab compute on dynamic supersteps |
Hardware-description-style register pattern
(BACK_EDGE) |
Verilog, VHDL | Synchronous-circuit register-on-back-edge semantics across tick boundaries | DagDB exposes it as a graph-database primitive, not a synthesis target |
| Graph-database query / declarative composition | Datalog, Cypher (Neo4j), SPARQL | Composable declarative queries over graph data | DagDB is Boolean-only and tick-evaluable; Datalog is full first-order logic with fixed-point semantics |
| MCP surface / tool exposure | Model Context Protocol (Anthropic, 2024) | LLM agent interaction with the database | Standard usage, no novelty claim |
Citations. - Pregel. Malewicz et al. (2010). Pregel: a system for large-scale graph processing. SIGMOD ’10. - GraphLab. Low et al. (2010, 2012). GraphLab: A new framework for parallel machine learning; and Distributed GraphLab. PVLDB. - Datalog. Ceri, Gottlob, Tanca (1989). What you always wanted to know about Datalog (and never dared to ask). IEEE TKDE. - Verilog / VHDL. IEEE Std 1364 / 1076. - MCP. Anthropic (2024). Model Context Protocol specification.
Label. VERIFIED prior art for every ingredient. The combination — LUT6-as-data + ranked-DAG + GPU-parallel evaluation + database-grade persistence + MCP surface, all on a single laptop — is what’s novel.
| Claim | Citation | Label |
|---|---|---|
| 7-coloring of hex squared graph | Molloy & Reed 2002; Salavatipour 2005 | VERIFIED prior art |
| Morton Z-curve, 2.11× speedup at 109 cells | Morton 1966 (algorithm); measurement ours | VERIFIED |
| Type II saturation breaks predator death spiral | Holling 1959 | VERIFIED prior art |
| Carlos Delta XOR + Zstandard at 50× lossless | Carlos Mateo, MIT-licensed | VERIFIED external |
| CFL clamp on dt-compression | Courant–Friedrichs–Lewy 1928 | VERIFIED prior art |
| DagDB ingredient combination novelty | FPGA + Pregel/GraphLab + Verilog + Datalog + MCP | VERIFIED prior-art map; combination is novel |
What’s not claimed. None of the math ingredients are novel. The contribution is the assembly: an ultra-scale spatial lattice engine on consumer Apple Silicon, with topological-cleanness proof (7-coloring) + cache-line locality (Morton) + Lotka–Volterra dynamics fixed (Type II) + lossless compression (Carlos Delta) + temporal-aliasing safety (CFL clamp). DagDB inherits the same dispatch + Morton + 7-coloring lineage from Savanna and adds LUT6-as-data + ranked-DAG semantics + the database layer. The combination matters; no single ingredient does.
../one_pager.md: “Novel
claims (any) — most ingredients have prior art” — VERIFIED prior-art
contextslides/direction9_theoretical_math.md
(open theoretical directions sitting on top of the same substrate)Stubbed by Pub on 2026-05-01 as PM coordinator. Drafter: ref. Draft 1, populated 2026-05-01. GH-link footer added 2026-05-02.
Inside the engine — UMA, Metal dispatch, and the lock-free wiring that holds it together.
Owner: dag (engine internals); pub (editorial pass)
Status: DRAFT 1 (2026-05-02) Tier: 2
(medium level / main directions) Calibration anchor:
../one_pager.md
One-slide architecture-detail layer underneath Direction #5 (DagDB). For the audience that wants the gear-level picture: how the pieces wire, what’s verified, what’s spec.
Required content: - Apple Silicon UMA + Metal compute — why unified memory matters for sparse graph traversal at scale (zero-copy GPU↔︎CPU) - Lock-free chromatic dispatch — 7 colour passes per tick, no atomics on entity update, atomic only on per-tick population census - Lazy mipmap tiles for trillion-cell scale — the I/O-death-spiral fix (Google-Earth-style lazy fetch, write-time mipmaps, NEVER assemble the 1 TB monolithic frame) - POSIX shared memory zero-copy — the dagdb-daemon ↔︎ MCP bridge ↔︎ Python adapter path - Dev/test/prod environment separation — Phase 1 shipped (data-layout + plist migration), Demerzel 6 supervisor for prod is SPEC
What this slide does NOT do: - Re-derive what’s in
one_pager.md Verified column — cite and pull labels - Claim
Demerzel 6 supervisor is built (it’s SPEC)
Headline. The gear-level picture underneath Direction #5. How the pieces wire on Apple Silicon, what’s verified today, what’s spec.
Apple Silicon UMA + Metal compute. One unified
memory pool shared by CPU and GPU. Sparse graph traversal at scale
benefits because there is no PCIe round-trip — the GPU kernel reads the
same bytes the CPU just wrote without an explicit copy. Engine buffers
(truth state, rank, LUT halves, neighbor table, edge weights) all live
as MTLBuffers with .shared storage mode; swift
code reads + writes the same memory the Metal shader operates on.
Verified — engine runs end-to-end this way.
Diagram lifted verbatim from
dagdb-engine/site/sql-architecture.html (slide
16).
Lock-free chromatic dispatch. Each tick fires seven colour passes (the 7-coloring of a hex grid means cells in the same colour class never share a 6-neighbour edge). Within one colour, node updates are independent — no atomics needed on the entity- update path. Atomic only on a per-tick population census. Inherits Savanna’s dispatch shape; the Boolean-LUT engine is leaner than the fluid-dynamics engine but uses the same scheduling. Verified.
POSIX shared memory zero-copy. The dagdb-daemon ↔︎
MCP bridge ↔︎ Python adapter path uses POSIX shm pages
(/tmp/dagdb_shm_file) for query result transport — Python’s
mmap + the daemon’s shared mapping share bytes without
socket-level serialization. Important when query results are large (full
secondary-index results, BFS frontiers, distance-metric outputs).
Verified — existing path in production.
Lazy mipmap tiles for trillion-cell scale. The path
from 10⁹ to 10¹¹: tiles paged to/from NVMe, two tiles resident at a time
(active + pre-fetch), cross-tile addressing via 24-bit-tile +
40-bit-local u64 global ID. Avoids the I/O-death-spiral of trying to
assemble a 1 TB monolithic frame in UMA — Google-Earth-style lazy-fetch
+ write-time mipmaps. 632-line spec at
docs/tiled-streaming.md. Step 4 scaffold landed (the
TiledGraphRouter actor, public surface). Steps 5–7 (live tile load, NVMe
streaming, cross-tile BFS continuation) pending.
Dev / test / prod environment separation. Phase 1
shipped 2026-05-01 — data layout migrated, plist updated. Phase 2
shipped 2026-05-02 — daemon reads
DAGDB_ENV ∈ {dev, test, prod}, derives
~/dag_databases/<env>/, fails loud on conflict.
Phases 3–4 (snapshot v5 env-origin stamp + socket rename to
/tmp/dagdb-prod.sock) batched on the env-split feature
branch. Phase 5 — -prod/ worktree pinned to release tag —
pending. Phases 1+2 SHIPPED, Phases 3–4 IN FLIGHT, full split
SPEC.
Demerzel 6 supervisor. Replaces the launchctl plist
supervision for the prod daemon. Three contracts: spawn (env var, args,
working dir, log path), health (STATUS poll, 200 ms readiness / 10 s
ongoing, degraded after 3 consecutive failures → markdown fallback +
drift queue), cleanup ladder (SHUTDOWN over socket → SIGTERM → SIGKILL
last resort). D6 Phase 1 shipped 2026-05-02 by Varpet —
supervisor core, hive_store.py with markdown-leader triple-write,
d6 CLI. Cutover from launchctl to D6 happens when my
env-split phases land. D6 PHASE 1 SHIPPED, cutover SPEC.
Triple-write pattern (with markdown as the leader). Hard requirement: the hive must keep working when DagDB is down. Every honey/journal mutation lands in markdown first (atomic file replace + F_FULLFSYNC, refuse the call on failure), then DagDB (best-effort, drift-queue on miss), then Postgres mirror via libpq from D6’s library (best-effort, drift-queue on miss). Drift-queue replay uses query-then-write client-side dedupe (daemon stays stateless on the command path). Designed jointly with Varpet; Phase 4c of honey-on-DagDB migration.
docs/tiled-streaming.md — 632-line specdocs/dev-test-prod-memo-2026-05-01.md — joint with
Varpet| Claim | Label |
|---|---|
| Apple Silicon UMA + Metal compute | VERIFIED — engine runs end-to-end via .shared
MTLBuffers |
| Lock-free chromatic dispatch (7 passes / tick) | VERIFIED |
| Atomics only on population census | VERIFIED |
| Lazy mipmap tiles for trillion-cell scale | SPEC — Step 4 scaffold (TiledGraphRouter actor) landed; Steps 5–7 pending |
| POSIX shm zero-copy | VERIFIED — daemon ↔︎ MCP bridge path in production |
| Dev/test/prod env separation | PHASE 1 SHIPPED 2026-05-01; PHASE 2 (DAGDB_ENV semantic) IN FLIGHT; PHASES 3–4 batched; PHASE 5 (-prod/ worktree) pending; full split SPEC |
| Demerzel 6 supervisor | D6 PHASE 1 SHIPPED 2026-05-02 by Varpet (supervisor core + hive_store.py + d6 CLI); cutover SPEC |
| Triple-write (markdown leader → DagDB → Postgres) | SPEC — designed jointly, lands as Phase 4c of honey-on-DagDB |
What’s not claimed. Tile-streaming for 10¹¹ on one M5 Max is spec, not built. The 632-line design exists; Step 4 scaffold landed; Steps 5–7 (live tile load, NVMe streaming, cross-tile BFS continuation) are pending. The 100B Savanna existence proof is what gives confidence the spec is reachable, not an inheritance claim.
Stubbed by Pub on 2026-05-01. Drafted by dag on 2026-05-02. Pub-screen pass before any external recording or distribution.
Source: github.com/norayr-m/dagdb-engine · tile-streaming spec · dev-test-prod memo
Mathematical directions the substrate brings within reach — directions of interest, not finished work.
Owner: ref (math); pub (editorial pass)
Status: DRAFT 1 — populated 2026-05-01
Tier: 2 (medium level / main directions)
Calibration anchor: ../one_pager.md
Directions of interest, not results. Three open theoretical threads Norayr is tracking. Surfacing them here is invitation to push back, name prior art, or name workloads that match. Nothing on this slide is a theorem; nothing is a published result. The hive engineering substrate (DagDB + the trio + the Savanna lineage) is what these directions could land on; they have not landed yet.
Spectral partitioning of hierarchical graphs on ranked spheres. Given a rooted, connected graph G = (V, E), embed it into ℝ3 by:
The resulting embedding is root-automorphism equivariant, localised (perturbations don’t cascade across independent branches), and separates rigid vs. flexible subtrees by area allocated to each cone.
Status. Four-page math paper drafted
(paper2_eigencone.tex in the eigencone subproject);
construction fully described, no code yet. Open directions named in the
paper:
The radial-coordinate-from-root construction in §1 is itself a stand-alone primitive: a graph carries a canonical decomposition into BFS shells, each shell treated as a discrete sphere with a spectral measure on it (the Fiedler-mass of each branch). Algebraic properties don’t depend on the subsequent Thomson placement: shell-by-shell mass conservation, root-action equivariance, local-perturbation locality.
Status. Open. Lives implicitly inside the eigencone write-up; not extracted as its own theorem about ranked graphs. Question for the audience. Is this exactly what graph-signal-processing already calls something else? It feels adjacent to spherical-harmonic decompositions on graphs (Hammond–Vandergheynst–Gribonval; Shuman et al.) but the BFS-shell-as-sphere discretisation is not standard as far as we know.
A family of distance metrics between subgraphs of a ranked DAG. The ranked DAG is the substrate (every DagDB instance is one); a subgraph is a node-set carved out of it; a distance metric returns a [0, 1] value. The direction of interest is what spectral and combinatorial distances are well-defined on subgraphs of bounded-fan-in ranked DAGs, beyond the standard set-overlap distances.
Engineered foothold (verified, in the substrate):
seven metrics shipped in DagDBDistance.swift:
| Metric | What it measures |
|---|---|
| Jaccard (nodes) | |A ∩ B|/|A ∪ B| on node sets |
| Jaccard (edges) | same on induced edges |
| Rank-profile L1 / L2 | DAG-shape signature via rank histograms |
| Node-type-profile L1 | type-distribution distance |
| Bounded GED | Jaccard symmetric-difference upper bound on edit distance |
| Weisfeiler–Lehman-1 | hash-histogram L1, neighbourhood-aware |
Open theoretical directions sitting on top of this engineering:
| Direction | Status | Notes |
|---|---|---|
| Eigencone constellations | OPEN — paper draft, no proof, no code | paper2_eigencone.tex is the artefact |
| Ranked spherical decomposition | OPEN — implicit in eigencone, not extracted | adjacent to graph-signal-processing literature |
| Ranked subgraph distances | OPEN — engineered foothold shipped, theory sparse, Norayr-prioritised | tissue-scale contact graphs as substrate; trio’s SpMV as computational layer |
What’s not claimed. None of these are results. They are the open theoretical directions Norayr is tracking. Surfacing them to a audience is invitation to push back, name prior art, or name workloads that match. The hive engineering substrate (DagDB + the trio + the Savanna lineage) is what these directions could land on; they have not landed yet.
Sources/DagDB/DagDBDistance.swift): https://github.com/norayr-m/dagdb-enginepaper2_eigencone.tex (private, not yet public;
pre-publication sharing on request)slides/hepatocyte_pass_2_spec.md (the hepatic-acinus
6-bound ranked DAG, 107
hepatocytes, 16 zonation ranks — this is the concrete substrate the
ranked-subgraph-distance question would live on)slides/direction7_applied_math.md (the
applied-math citations + lineage that this slide’s open directions sit
on top of)Stubbed by Pub on 2026-05-01 as PM coordinator. Drafter: ref. Draft 1, populated 2026-05-01. GH-link footer added 2026-05-02.
Tier 1 narrative, side material, full inventory walk, and the live-Q&A handoff. Here for those who want depth or want to review.
Calibration: every line below maps to a label in the project’s calibration baseline (verified / spec / overclaim risk). Numbers from one M5 Max laptop. Amateur engineering project; no competitive claims; errors likely. Engine source is the source of truth; if anything below contradicts the code, the code wins.
One M5 Max laptop. Three months. Three things landed end-to-end.
Savanna Engine — ultra-scale spatial lattice on Apple Silicon. 100 billion cells over 9 hours on one M5 Max, 15.8 GCUPS peak, 500 GB state file at the M5 thermal ceiling. VERIFIED as an existence proof for what consumer hardware can actually do under careful dispatch + Morton-Z-curve memory layout + 7-coloring parallel-safe scheduling.
DagDB engine — Boolean-circuit-as-database. A graph database where every node holds a 64-bit truth table (a LUT6 — the same primitive at the heart of an FPGA), wired into a ranked acyclic graph evaluated in parallel on the GPU each tick. BACK_EDGE primitive (synchronous-circuit register pattern) verified by AC-3 constraint-propagation round-trip against a Python reference: per-tick equality across 21 register nodes, converges in 2 synchronous ticks. Snapshot v4 with WAL replay, multi-version reads, atomic save discipline; bitwise LUT composition at runtime; honey-on-DagDB lossless round-trip probe. VERIFIED.
Bio digital twin substrate. Graph-CA-on-topology framework that takes the Savanna throughput and the DagDB substrate and targets tissue-scale biological models. Hepatocyte 10⁷ Pass 2 spatial-adjacency schema designed and cross-checked. Integration path with allosteric-pathway protein work in active development. MIXED — the substrate is verified at the throughput tier; specific tissue twins (liver, brain cortical column) are spec, not built.
Underneath the three engineering landings: a discipline of
labelling claims. Every artefact in this presentation maps to
one of three labels: verified (compiled, tested,
measured), spec (designed, not built), or
overclaim risk (something a casual reader might infer
that isn’t actually true). The calibration anchor
(one_pager.md) names the specific overclaim risks for this
project — for example, “DagDB is a thinking engine” is overclaim (it’s a
fast specialised substrate, not a general reasoner); “the Tokyo CA
produces a slime-mold network” is overclaim (it produces a Voronoi
tessellation; the substrate has no fluid dynamics).
The honest split is the deliverable. The numbers are real; the limits are named.
Anyone who has not seen the work before should walk away from these five beats with: (a) what was built, (b) at what scale, (c) what’s real vs spec, and (d) the labelling discipline that carries through the rest of the deck.
Tier 2 expands each direction; Tier 3 is the clickable inventory. Numbers above are the substantive ones; everything else is texture.
Honest pushback on claim calibration. HPC specialists see more of this work than we do; if any label above is too generous in either direction, we want the redirect. Pointers to prior art are also welcome — the “novel combination, prior-art ingredients” framing only works if we have the ingredients named accurately.
Slot owner: tut (explanatory framing). Co-owner: pub
(calibration discipline policy). Calibration source:
one_pager.md. If a claim above contradicts the engine
source, the engine source wins.
Source repos: Savanna · DagDB · drt-generator · drt-scanner · drt-cell-simulator · isomorphic-walk. Profile: github.com/norayr-m.
Calibration: this slide describes a development discipline, not a library or an engine. The substrate is one human operator using multiple AI agents in parallel under tight per-agent specialisation and explicit calibration policy. What’s claimed below is a way of working; what’s claimed about output is governed by the project’s calibration baseline (
one_pager.md).
A multi-agent development discipline running on one M5 Max laptop. Different specialised agents own different lanes — engine, math, gatekeeping, infrastructure, life-admin — coordinated through file-based asynchronous messages and a small set of structural conventions. The substrate is plain code (Swift, Python, Metal) and plain protocols (markdown, JSON, git, launchctl, Metal compute). The discipline is what keeps the output honest.
Per-agent git worktrees. Each specialised agent develops in a dedicated git worktree on a dedicated branch. No shared HEAD races; agents do not stomp each other’s edits. A real engineering problem (we hit a shared-HEAD race four times in two hours one evening before adopting this convention) with a structural fix.
Asynchronous file-based comms (drops).
Coordination between agents flows through markdown drops in a shared
directory. Each drop has a fixed naming convention
(YYYY-MM-DD_<from>-to-<to>_<subject>_<AGENT>_v1.md)
and a single-recipient or list-of-recipients header. Receipts close
loops; cross-references stay grep-able indefinitely.
Calibration discipline. Every public-facing artefact maps each of its claims to one of three labels: verified (compiled, tested, measured), spec (designed, not built), or overclaim risk (something a casual reader might infer that isn’t actually true). The labels are explicit on the artefact itself, not buried in a separate honesty-disclaimer page. A gatekeeping-role agent screens every public artefact against this discipline before it leaves the laptop.
Retraction protocol. When a claim that previously shipped turns out to be wrong, the retraction is named on the relevant page — not silently rewritten. The canonical case is a norm-growth inequality from earlier work that was found to be tautological under closer inspection; that retraction is now the named precedent for how subsequent claims handle being shown to be wrong. The discipline is “retract loudly, in-place.”
Persistent-memory architecture per agent. Each agent maintains a local “honey” file of learned facts and a journal of session work. A common honey is shared across all agents. Memory is plain markdown; it survives session restarts and is grep-able by the operator. No magic.
The throughput numbers in the rest of this deck (Savanna 100 billion cells, DagDB BACK_EDGE/AC-3 verified, etc.) come from work organised under this discipline. The discipline is not load-bearing for the results — the engines are real, the tests pass, the numbers are measurable on independent hardware. The discipline is load-bearing for the honesty of the labels around the results: distinguishing what was actually measured from what was designed but not built, and calling out what a casual reader might over-infer.
Two specific consequences:
one_pager.md)
carries an explicit “what’s overclaim risk” section listing four claims
a reasonable reader could infer that the work does not support. That
section was not added defensively after the fact — it was written in the
same draft as the verified-and-spec sections, by the same agent, under
the same screen.This slide describes a way of working, not a measured artefact. If the audience wants to see the discipline applied: every label in this deck (verified / spec / overclaim) and the named-retraction references on relevant pages are the in-deck evidence. The machinery underneath (boot validators, screen checks, honey files, ring registry) lives in a forthcoming companion architecture document that walks the substrate in detail.
Slot owner: tut (explanatory framing). Co-owner: pub
(calibration discipline policy + screen). Coordinated with: vpt
(architecture document forthcoming). Calibration source:
one_pager.md. Substrate description: companion
hive-architecture document, currently in v0.1 review.
Calibration: substrate-level throughput is VERIFIED at the indicated scales. Specific tissue twins (liver lobule, full liver, brain cortical column) are SPEC — designed, schemas drafted, not built. The framing here is “what we are building toward,” with the verified-vs-spec split made explicit on every claim. Engine source remains source of truth.
A real-time digital twin of biological organs at single-cell or near-single-cell resolution, running on one consumer M5 Max laptop. The substrate is the graph-CA-on-topology framework underneath DagDB and Savanna: nodes are cells (or cell aggregates), edges are spatial-adjacency or signal-flow, ranks encode the directional gradient (blood-flow, signal-cascade, periportal-to-pericentral).
The deliverable, when complete, is the ability to:
Three classes of biological question that today’s tools handle poorly at tissue scale:
These are real biological questions with real clinical relevance. Whether the substrate proposed here actually answers them depends on validation work that is not part of this work’s claims; the substrate is one ingredient.
| Tissue | Cell count | Status |
|---|---|---|
| Liver lobule (single functional unit) | 10⁶ hepatocytes | SPEC — schema designed (HepaticZonationPolicy, 16 zonation ranks); not built |
| Full liver | 10⁸ hepatocytes (~10⁷ for the Pass 2 schema) | SPEC — Pass 2 schema agreed with the protein-pathway lane; not built |
| Brain cortical column | 10⁵ neurons | SPEC — concept stage |
The substrate-level throughput is verified at the relevant order of magnitude (the Savanna 10¹¹-cell result comfortably contains a 10⁷ hepatocyte simulation in raw cell-count terms). What is not yet verified is that the substrate, configured with the specific HepaticZonationPolicy and the specific damage-cascade LUT logic, will reproduce biology faithfully. That is validation work, not substrate work, and it has not been done.
The honest framing: substrate verified, biology validation pending.
Slot owner: tut (explanatory framing). Technical
co-owner: fold (hepatocyte schema, isomorphic-walk integration).
Calibration source: one_pager.md. Engine source
authoritative. Specific tick-rate numbers cited at substrate-level
throughput gate against the Savanna result and the DagDB measured ticks
at 1M nodes (~2 ms/tick at one million nodes, M5 Max,
verified).
Source repos: substrate engines — DagDB, Savanna; bio digital twin prototype — drt-cell-simulator; allosteric protein lane — isomorphic-walk.
Calibration: this slide reports a SPEC, not a built artefact. The underlying graph engine is verified at the 10⁹ node tier; the hepatocyte-specific deployment described below is queued, not yet built. The schema was cross-checked with the engine sibling on 2026-04-29.
A graph substrate for representing a human liver at single-hepatocyte resolution. 10⁷ hepatocyte nodes arranged as a spatial-adjacency graph: each cell connects to its 4–6 nearest neighbours in the plate- and-sinusoid morphology of the liver acinus. The connectivity pattern is natively bounded at 6 incoming edges per node, which fits the DagDB engine’s 6-bound DAG invariant without virtual node splitting.
Rank policy: HepaticZonationPolicy —
rank(cell) = quantize(distance_from_portal_triad, n_levels=16).
This converts the biologically real periportal → pericentral functional
gradient (zone 1 → zone 3, characterised since the 1970s in the
histology literature) into the engine’s monotone rank ordering. Edge
direction becomes “blood-flow direction” (portal → central), which is
also the direction along which hormones, nutrients, and damage signals
propagate biologically.
Ticks: quasi-static graph by default + opt-in LUT6 propagation for damage / signal cascade queries.
Three classes of question that matter biologically and that today’s tools handle poorly at tissue scale:
The 10⁷ scale is meaningful: a human liver lobule has roughly 10⁵ hepatocytes; a complete liver has ~10¹¹. 10⁷ is a few hundred lobules — small enough to fit single-engine on an M5 Max laptop (≈420 MB at 42 B/node, 0.3% of UMA), large enough to capture multi-lobule spatial gradients that single-lobule models miss.
DagDB engine — 6-bounded ranked DAG with LUT6 gates, currently verified at the 10⁹ node tier (single engine, no tiling). 10⁷ hepatocyte target sits at the single-engine, non-tiled tier per the engine spec — does not depend on tile-streaming landing first. Schema cross-checked with the engine sibling on 2026-04-29: spatial-adjacency layer + HepaticZonationPolicy ranks + quasi- static-with-opt-in-propagation tick semantics.
Nodes (10^7):
type: hepatocyte
rank: HepaticZonationPolicy (0 = portal, 15 = central)
attrs:
coords: (i, j, k) — grid position
zone: 1 / 2 / 3 (classical zonation)
truth: 0 healthy (default)
1 damaged
2 apoptotic
(used only during propagation ticks)
Edges (~5 × 10^7):
type: spatial-adjacency (hepatocyte ↔ hepatocyte plate contact)
policy: source.rank > dst.rank (blood-flow direction)
6-bound: holds natively (~0.1% loss at corner cells)
Rank levels: 16 (3 zones × ~5 sub-zones each)
Tick: quasi-static; opt-in LUT6 propagation for damage cascade
each tick step ≈ hour-scale physical time
Out of scope for Pass 2:
signaling-pathway edges (degree 10²–10³, needs virtual splitting)
sinusoid hub nodes (degree 10³–10⁴, different node type)
metabolic state simulation (needs activation channel + Pass 3 design)../one_pager.md: “Hepatocyte 10⁷
Pass 2 schema: spatial-adjacency layer with HepaticZonationPolicy ranks
(16 levels, periportal → pericentral). Schema agreed with Fold; not
built.” — SPEC tierslides/direction6_proteins.md (the
protein-pathway prediction lane that motivates the tissue-scale
substrate question)This slide is part of the the work deck. Calibration discipline locked: verified vs spec vs overclaim-risk three-column split. This artefact is in the spec column.
Calibration: numbers in this slide come from the verified tier of
one_pager.md— measured on M5 Max (128 GB unified memory), 10-run validated, <1 % standard deviation. Any number outside that anchor is flagged. Existence proof for the scale path inherited from the Savanna predecessor (100 B cells over 9 hours, same hardware).
DagDB runs on Apple Metal GPU compute with the entire graph state resident in unified memory — a single physical RAM addressed by both CPU and GPU with zero copy across the boundary. Three direct consequences:
cudaMemcpy, no DMA-in / DMA-out, no
double- buffering. The 23 MB of per-cell state at 1 M nodes lives in one
place.BACK_EDGE two-phase snapshot/commit runs on the CPU at
sub-ms per pass for 1 M back-edges (untested at scale, plausible but not
yet measured). Deferred GPU-side latch kernel is on the shelf.Caveat: M5 Max specifically. Smaller M-series chips have less unified memory and thermal headroom; the scaling table below is for M5 Max only.
| Quantity | Value | Notes |
|---|---|---|
| Grid | 1 048 576 nodes (1024 × 1024 hex) | |
| State per node | 23 MB total / 1 M cells = ~23 B/node | 5 channels + 4 scent fields |
| Per-tick compute | ~2 ms | 13 kernel dispatches |
| Throughput | ~500 k ticks/s GPU only | excludes display, recorder |
| Dispatch overhead | 5 % GPU utilization | rest waiting on vsync / CPU |
| Memory bandwidth | 29 GB/s | observed during tick |
| 10-run σ | < 1 % | reproducible |
For the Tokyo CA workload at 200×200 (smaller scale, more nodes per cell): ~few ms per tick — same envelope as the 1 M-cell result above with different per-cell state shape.
DagDB shares dispatch + Morton + 7-coloring + halo file format with the predecessor Savanna engine. Savanna ran:
| Grid | Wall time | Throughput | State |
|---|---|---|---|
| 1 M cells | sub-second | 1 600 tps | 23 MB |
| 16 M cells | ~13 sec | 77 tps | 367 MB |
| 100 B cells | 9 hours | ~3 tps* | 500 GB |
*100 B run hit M5 thermal ceiling; 9-hour wall reflects sustained throttled state, not headroom. Same dispatch machinery, different per-cell state.
This is existence-proof for the substrate’s scale path. DagDB ran the same machinery; the per-cell state happens to be Boolean LUT6 + ranked DAG metadata instead of Savanna’s fluid + scent + entity tables. The throughput envelope is determined by the engine machinery, not the workload semantics.
docs/tiled-streaming.md.
Steps 1-2 + 4 shipped (TileHalo, GlobalNodeID, TiledGraphRouter
scaffold). Steps 5-7 (live tile load, NVMe streaming, cross-tile BFS)
pending.BACK_EDGE: the CPU-side two-phase latch is fine at 1 M
back-edges; for 10¹² this becomes the bottleneck and a GPU-side
equivalent is shelved as a known next-step optimization.The numbers above are real, on real hardware, with real load. Three things to flag before the audience pulls them:
dagdb-engine/docs/tiled-streaming.md../one_pager.md: “~2 ms per tick at
1 M nodes on M5 Max” — VERIFIED tierslides/direction4_savanna.md (the 100
B / 9 h Savanna run that anchors the scaling table)UMA performance slide: IN PROGRESS
(status board at README.md). - [x] This slide drafted - [x]
Why-UMA SVG inlined (pandoc indent fix landed 18:51) - [x] GH-link
footer added - [ ] Verify ~2 ms and ~500 k tps
on current main (re-run benchmark) - [ ] Pub-screen pass
bzz. — slvr.
Calibration: this slide reports a VERIFIED build. The 200×200 Tokyo Greenberg-Hastings cellular automaton runs end-to-end on the DagDB substrate; convergence + dashboard + per-tick recording all measurable on current main. Honest framing guard: this is NOT a slime-mold network — that’s an explicit overclaim flagged in
one_pager.md. The result is a Voronoi tessellation between food sources, produced by wave-front collision. Seedemos/tokyo_ca.md(consolidated demo run plan + technical encoding).
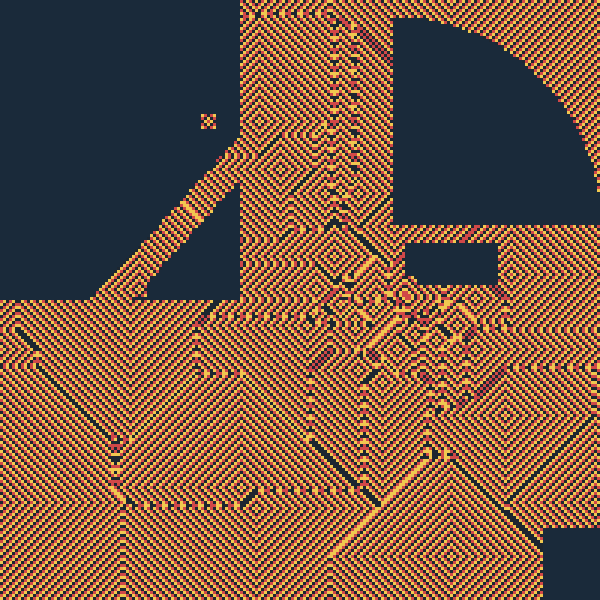
A 200×200 hex-grid cellular automaton, encoded as a ranked DagDB DAG with the Greenberg-Hastings 3-state excitable rule:
Food cells (36 Tokyo-region city positions) re-trigger excitation periodically, producing radial wave fronts. Wave-front collisions between adjacent food cells trace out the Voronoi tessellation boundary — the locus of cells equidistant (in hop count) from two food sources.
Per cell on the substrate: - 11 logical nodes per cell × 40 000 cells
≈ 440 000 DagDB nodes total - 3 phase-bit register
nodes per cell, latched by BACK_EDGE - 8 combinational LUT6
nodes per cell for neighbour excitation, in-range detection, next-state
logic - All evaluations rank-stratified for parallel-safe dispatch
Convergence to the steady-state Voronoi pattern at ~tick 160.
Three substrate properties this demo proves visibly:
BACK_EDGE works at scale. 440 K nodes
with 3 register nodes per cell, all latching cleanly tick after tick.
The synchronous-circuit register pattern is the load-bearing primitive
that makes recurrent dynamics on a 6-bounded DAG possible.This is the load-bearing framing for the audience. Three things to say up front in the live demo:
BACK_EDGE register latching, on a
GPU substrate with WAL + multi-version reads — same ingredients as a
real-time database, applied to a CA workload.| Quantity | Value | Source |
|---|---|---|
| Grid | 200 × 200 hex cells | terrain spec |
| Land cells | 28 038 | inert mask filtered |
| Inert cells | 11 962 | water + Pacific + mountains |
| DagDB nodes | ~440 000 | 11 nodes/cell × 40 000 cells |
BACK_EDGEs |
~120 000 | 3 register nodes × 40 000 cells |
| Per-tick wall | ~24 ms on M5 Max | with full per-tick dashboard recording; pure compute is faster |
| Convergence | 160 ticks ✓ | swift test TokyoCATests/testTokyoSolve200x200 re-run
2026-05-01 |
| Total wall | 4.6 s for 190 ticks | live demo runs in real time |
| 7-color groups | 7 | hex lattice distance-2 chromatic number (Molloy & Salavatipour 2005) |
Primary:
assets/tokyo_ca_voronoi_t160.png (200 × 200 → 600 × 600
upscaled, NEAREST). Yellow-orange checkerboard texture is the
oscillating excitable medium; bright yellow ridges across the field are
wave-front collision lines = Voronoi edges between food cells. Central
diamond pattern = waves converging from the Tokyo cluster. Dark-blue
regions are inert mask (Pacific, Tokyo Bay, mountains).
Time series (also in assets/):
tokyo_ca_t030.png, tokyo_ca_t060.png,
tokyo_ca_t100.png — show convergence progression for the
slide’s mid-presentation animation.
Greenberg, J.M. and Hastings, S.P. (1978). "Spatial patterns for
discrete models of diffusion in excitable media." SIAM J. Appl.
Math. 34(3), 515-523.
— origin of the 3-state excitable CA rule we run.
Tolmachev, D. and Adamatzky, A. (1996). "Chemical processor for
computation of Voronoi diagram." Adv. Mater. Optics Electron. 6(4),
191-196.
— wave-front collision on excitable media computes Voronoi as a
by-product. Direct precedent for what Tokyo demo shows.
Adamatzky, A. (2010). Game of Life Cellular Automata. Springer.
— Ch. 17 covers maze-solving / Voronoi-computing CA class. Our
Tokyo CA is in this lineage, NOT Tero 2010.
Murray, J.D. (2003). Mathematical Biology II. Springer, 3rd ed.
Chs. 1, 8-10. — accessible textbook treatment of excitable media
for the architect-grade audience.
Molloy, M. and Salavatipour, M.R. (2005). "A bound on the chromatic
number of the square of a planar graph." J. Combin. Theory B 94(2),
189-213. — distance-2 chromatic number 7 of the squared hex lattice;
underwrites the 7-coloring parallel dispatch.(Pulled from one_pager.md for completeness.)
“Tokyo CA runs slime mold on the substrate” — wrong. Substrate has no fluid dynamics; engine source explicitly says “No scent diffusion. No fluid dynamics.” Engine docstring is the source of truth. What the Tokyo demo actually shows is a Greenberg-Hastings excitable cellular automaton producing a Voronoi tessellation between food sources, NOT a slime-mold network. Real CA, real convergence, real visualization — but not what Tero 2010 produced biologically.
The mold_walk side demo sits next to this slide on
the deck and shows what actually trying for slime mold looks
like — on a different substrate, explicitly labelled as Metal-side (see
slides/mold_walk_side_demo.md).
dagdb/Tests/DagDBTests/TokyoCATests.swift
(testTokyoSolve200x200 — convergence at tick 160, 4.6 s
wall)dagdb/examples/tokyo_ca/build_dashboard_data.py +
build_dashboard_html.py~/000_AI_Work/0_Projects/0_Active_MoldWalk/ (referenced by
side demo, not the DagDB run)Tokyo CA live demo: IN PROGRESS (status
board at README.md). - [x] Demo run plan + technical
encoding consolidated (demos/tokyo_ca.md) - [x] This slide
drafted - [x] UMA performance slide drafted
(uma_performance.md) - [x] Visual asset inlined
(tokyo_ca_voronoi_t160.png) - [x] GH-link footer added - [
] Side mold_walk slide (tomorrow afternoon) - [ ] Convergence GIF
capture under assets/ - [ ] Pub-screen pass on the GH
footer addition
bzz. — slvr.
Calibration: this is the clickable detail behind every direction in Layers 1 and 2. White papers, live demos, repositories, demo scripts, math papers, media. Maintained as new work lands; post-recording this section migrates to the GitHub Profile README as the live wiki end-state.
Six categories. Every row links to a real public artefact.
The narration walks the structure; this slide carries the URLs you click.
| Paper | URL |
|---|---|
| Savanna Engine — Architecture, Build Approach, Overcomes | norayr-m.github.io/savanna-engine/whitepaper.html |
| Generator + Cell Simulator + Scanner — sparse mat-vec trio | norayr-m.github.io/drt-generator/whitepaper.html |
| DRT Scanner — inversion-fidelity diagnostic | norayr-m.github.io/drt-scanner/whitepaper.html |
| DRT Cell Simulator | norayr-m.github.io/drt-cell-simulator/whitepaper.html |
Five more queued (Decoder, Pipeline, Interstellar, Orbital, Encoder Brain). Engine substrate paper held for explicit trigger.
DagDB family: engine landing · SQL architecture · grid demo · podcast · interview · wiki
Savanna family: interactive presentation · 7-coloring · 1B cell playback · Morton charts · about
Bio digital twin: brain-tile prototype
DRT-X canonical 5-collection: Composer · Scanner · Pipeline · Cell Simulator · Bio D Twin · plus Generator presentation deck
DRT-X standalones: Decoder of the Encoder · Interstellar Harmony · Orbital Solver · Encoder Brain
Other public: Three Basket Backup · Ternary Lattice · Isomorphic Walk viewers
Twenty-five browser-runnable demos at recording time.
Thirteen active public repositories, plus the Profile.
dagdb-engine · savanna-engine · isomorphic-walk · drt-generator · drt-scanner · drt-cell-simulator · decoder-of-the-encoder · DRT-Encoder-Brain · drt-pipeline · drt-orbital-solver · interstellar-harmony · three-basket-backup · ternary-lattice
Profile: github.com/norayr-m
Five scripts in flight: demo order narrative, BACK_EDGE / AC-3 live demo, Tokyo CA on DagDB, honey-on-DagDB, tile-streaming spec walkthrough.
Distributed Reconstruction Theorem v0.1 — in preparation.
Ternary firewall topological regularization (CVPDE) — in preparation, separate venue, not the focus of this work.
YouTube: Savanna v0.6 · Swift Infection · Swift Haboob · 100B Cell test
Podcasts: DagDB Podcast (Carlos persona) · DagDB Dialogue (same site) · DRT Generator presentation deck narration (embedded)
Interviews: DagDB Podcast Interview
This inventory updates in the same push that lands the work. No separate sync step. Eight-category screen runs every time. Post-recording the structure migrates to the Profile README as the live wiki end-state.
Slot owner: tut (explanatory framing on the inventory walk).
Source-of-truth: tier3_inventory.md. Calibration
source: one_pager.md.
Active public repos and Profile: github.com/norayr-m.
Live segment. Each specialist is active in their own tmux session during the Zoom and answers in their own voice via the live audio pipeline.
Direct your question to a specific specialist by name. Norayr relays. The specialist answers on the fly through the live voice pipeline — their actual voice, their actual lane, real-time.
The deck’s multi-voice narration is the rehearsal; the Q&A is the live demonstration.
| Specialist | Lane | Best for questions about |
|---|---|---|
| Tut | Explanatory framing, math foundations | The hive itself, the bio digital twin engineering goal, why-this-matters questions, math-foundation context |
| Slvr | Performance scout, the substrate at scale | Tokyo CA, UMA performance, the Savanna direction, perf measurements, scaling ceiling |
| Dag | Engine, DagDB internals, architecture | BACK_EDGE / AC-3, snapshot v4, MVCC, COMPOSE LUT, tile-streaming spec, gear-level questions |
| Fold | Proteins, allosteric pathway prediction, hepatocyte schema | Isomorphic Walk, HIV-1 capsid scaling, allosteric cohort, hepatocyte 10⁷ schema |
| Ref | Applied math, citations + lineage, theoretical directions | Molloy–Salavatipour 7-coloring, Morton Z-curve, Holling Type II, Carlos Delta, eigencone / starcone / ranked subgraph distances |
| Emma | Through-line spine, presentation flow | Deck navigation, layer structure, recording conventions |
Name the specialist + the question. Norayr passes it to the specialist’s session; their answer lands in audio within seconds.
If you’re unsure which specialist owns a question, name the topic and Norayr routes.
one_pager.md).
If a specialist doesn’t have verified-or-spec material on a topic, the
answer will say so explicitly.The discipline is the same as the deck’s: show what the work IS — name what it isn’t only when directly asked.
This Q&A operates under the same calibration discipline as the
deck. Every answer maps to verified, spec, or overclaim-risk per
one_pager.md. Specialists self-label their answers at the
appropriate tier.
If a specialist hits a question outside their lane, they hand off cleanly: “That’s [other-specialist]’s lane — [name], over to you?”
Slot owner: tut (explanatory framing on the section).
Live-event ownership: pub (production), varpet (tmux
infrastructure), each specialist (their own answers).
Calibration source: one_pager.md. Engine source
authoritative.
Active public repos — DagDB · Savanna · Isomorphic Walk · drt-generator · drt-scanner · drt-cell-simulator · drt-pipeline · drt-orbital-solver · decoder-of-the-encoder · DRT-Encoder-Brain · interstellar-harmony · three-basket-backup · ternary-lattice. Profile: github.com/norayr-m.